|
“Exactly what are the genetic differences between my sequenced organism and another related strain?” One common question coming out of this has been to ask: In recent years there has been an explosion of whole-genome sequencing projects. MacVector to the rescue! MacVector’s Compare Genomes By Feature… tool lets you see the differences between two annotated genomes in fine detail. The algorithm takes every annotated feature from the source genome and looks for the presence of that feature in the comparison genome based on sequence similarity. 
CDS features are even translated so that the predicted amino acid sequences are compared. The results are then tabulated to show identical, closely related, and weakly related features in separate tabs, with additional tabs for features that are completely missing and a “details” tab that shows the low-level alignment details for any matching pair of features. Hot-links in the result tabs let you quickly scroll the parent sequences to any individual feature of interest. How to compare a pair of genomesĬompares two related annotated genomes (or smaller sequences) to identify and list, in spreadsheet form, identical, similar and weakly similar features along with missing features. Open the pair of sequences you want to compare.Show amino acids snapgene viewer download#.Show amino acids snapgene viewer update#.Show amino acids snapgene viewer how to#.Getting Started with MacVector: An overview of primer design workflows in MacVector.Melissa Caimano on HOW DO I video guides to common molecular biology workflows.admin on HOW DO I video guides to common molecular biology workflows.mariam abdelmalak on Major release details – Summary.Brian on Designing primers and documenting In-Fusion Cloning with MacVector.Chris on Designing primers and documenting In-Fusion Cloning with MacVector.How to call heterozygotes in trace files or Assembly Projects.MacVectorTip: How to Customize the Toolbars of MacVector windows.MacVectorTip: Selecting the sequence from a single restriction enzyme site to the end of a linear sequence.Sequence Assembly: What can Assembler do for my lab?.You can learn more about the different alignment options in these three blog posts, Alignments in MacVector, Aligning Trace files, and Aligning primers to a Reference sequence. This lets you define tails for your primers and you can even select a pair of primers in the graphical result window and Edit | Copy will copy the predicted amplification product, complete with any engineered mismatches and overhanging tails. 
A relatively new addition to MacVector, but you can use this to align a collection of primers to a test sequence. Handles fasta and fastq files so that you can use this to pick out reads from a huge database that match your sample sequence. Like BLAST, but for your own personal sequences. But still a great way of getting a quick idea of which sequences your sample is related to. MacVector’s implementation of the standard NCBI BLAST search. 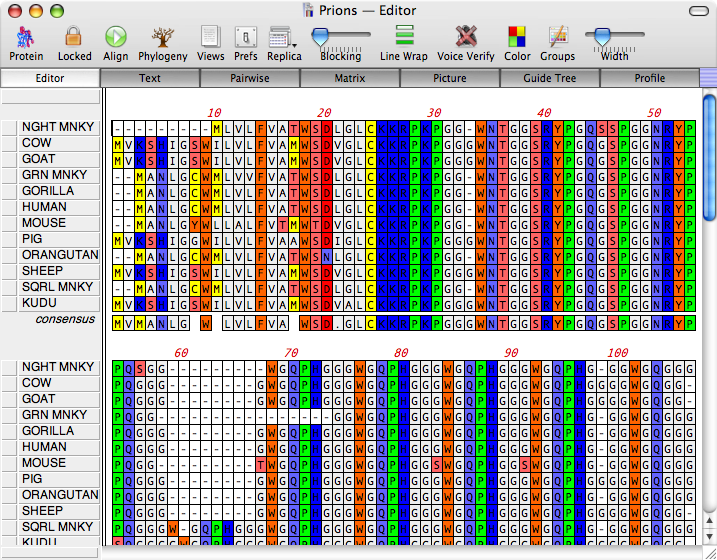
This is also a good way to see the extent of local alignments between two sequences. You can use this to quickly check that two sequences do have some similarity, but you can also use it to compare two genomes and see were the identities, duplications and inversions are located. This is a great and underrated function for quickly comparing 2 sequences to get an overview of the similarity between them.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |


 RSS Feed
RSS Feed
